NMM: Algorithms for Proteomics
Project focus: Protein feature determination from subsampled peptide sequences
Motivation:
Existing proteomic analyses depend heavily on use of tandem mass spectroscopy (MS/MS) to identify peptide backbone sequences. This technique currently requires relatively high protein copy number (100s of thousands to millions of copies of the same protein), making it inappropriate for a number of applications including in situ proteomics. At the same time, in situ protein sequencing would be of great benefit for understanding cell function, particularly in the nervous system or other organ systems with very dynamic protein regulation networks.
Summary:
I began this project envisioning a direct sequencing methodology with a number of possible physical instantiations. Consider e.g. starting with an unknown protein which typically consists of ~1500 amino acid (AA) residues or less, linearly ordered, broken up into smaller linear sequences using enzymes and/or reagents which cut the protein at specific sites (e.g. Cyanogen Bromide cuts the peptide after each Methionine residue...usually). Further, consider being able to uniquely label a minority of the 22 residues of the protein, and assume that a complete library of all proteins is available (i.e. our unknown protein is one of the library elements). (N.B. This turns out to be relatively close to reality: NCBI has compiled ~ 30k known human proteins, which is near the expected number). The goal, then, is to reconstruct the unknown protein with as few AA fragments as possible, with as much fidelity as possible.
(It should be noted that even constrained to existing MS/MS instrumentation, it appears that reconstruction of protein sequence information can benefit from a compressive sensing approach - see below!).
Along the way to this goal, a few optimization problems surface:
This set of optimization questions suggests the need for a quantitative measurement of the "uniqueness" or quality of the resulting tagged fragment library. The ideal fragment library would have a very low bit-score, that is, each fragment would be as dissimilar as possible from every other fragment. (BLAST is a very analagous bit-score, and used frequently in proteomics).
First Approaches:
Initially I created simulated cleaves of a subset of a ~20k protein FASTA database (cleaned of duplicates) using Cyanogen Bromide, a common reagent, and explored approximate string matching to see what fraction of proteins could be identified with n unique tags (where n is small). The simulation then gives a protein library recast in terms of these unique tags, e.g. K,Y,C and wildcard X for all other proteins.
This problem is highly idealized (for instance, there is no noise) and as such, the result is relatively uninteresting: with just three tags (K,Y,C) and one fortuitously chosen reagent (CyBr), 99.4% of proteins were idnetified uniquely. 7 proteins were lost due to choice of tags, and 110 due to choice of reagent. In this framework, ~80% of proteins are uniquely identifiable from a single AA fragment!
This result is highly idealized and breaks down as soon as noise is introduced into our signal thorugh protein tagging errors (e.g. tags not attaching due to steric hindrance, etc), cleavage errors (CyBr is not ideal - some cleaves do not occur where expected, others occur where unexpected), and/or read errors. A more noise-immune approach is necessary.
Compressed Sensing Framework:
Looking at this problem another way, sparsely sampled peptide fragments and the need for robustness in the presence of noise points - tentatively - to a compressive sensing approach (tentatively because we need to confirm that the sampling problem meets a few criteria, notably the Restricted Isometry Property - for more details, Candes and Wakin wrote a nice tutorial on CS, freely available here). We may be able to formulate a basis set representing all proteins as combinations of AA fragments such that using this basis set, we could represent a particular protein sample (or set of protein samples, if we wish to minimize purification steps) as:
Sample = phi*F*p, where:
I found the biggest challenge in framing this problem as a CS problem was to develop the matrix notation for a non-numerical data set (e.g. how to represent each protein fragment of abstract AA sequences numerically?).
Working with Mike Henninger and input from the Camera Culture group, we recast this problem once again, beginning with a detection problem for tandem mass spec (MS/MS) data. Rather than working with fragment sequences then, we only see AA fragment weights directly. This is a convenient place to start thinking about the actual dictionary optimization problem and CS mechanics.
Compressive Sensing for MS/MS:
Working from the abstraction of the protein library, it is straightforward to simulate the MS/MS readout: for each fragment, AA molecular weights are assigned to each residue, and a total molecular weight is computed for each fragment. The library of proteins then becomes a library of mass profiles.
(In this framework the G = phi*F matrix still serves the same role but the "tags" are implicit by protein mass. Each column of the matrix is a single mass spec-like mass profile)
Dealing with noise:
To add further characteristics of MS/MS to the simulation, bin size for mass profile is an adjustable parameter - defaulting to 7 Daltons as the resolution of mass peak width based on (Perkins 1999).
As noted already, reagents are not 100% reliable in terms of cut site specificity and completeness, thus any viable algorithm must be able to deal with missing peaks in the mass spec profile. As a simulation, we add a noise factor to create a noise mask by randomly removing peaks from the phi*F matrix, using error rates from (Perkins 1999).
Dictionary optimization:
Before going too far into optimizing the selection of reagents and tags to use, we need to define a metric for optimality. Effectively we're defining a projection from high dimensional space (sample) to low dimensional (measurement), which should be as orthogonal as possible. Along a similar approach to Marwah 2013 we characterize the optimality as a minimization on:
Again, G = phi*F matrix from above, which is the mass profile of proteins resulting from cleaving with a specific reagent. Further, we are normalizing by the number of elements in the matrix to handle varying dictionary sizes.
In the ideal case, G_transpose*G is the identity matrix, so our GOF would be zero.
Evaluating a few test cases on a subset of proteins for computational simplicity, with this metric we find:
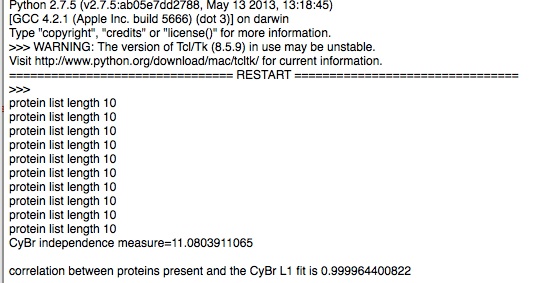

As seen, CyBr exhibits lower value for GOF and should therefore be a better reagent than Trypsin for uniquely identifying proteins. This holds in reality.
Implementing Compressive Sensing:
We now have our underdetermined matrix problem, sample = phi*F*p (or eqivalently using above notation, sample = G*p), and wish to solve it using compressive sensing. The approach implemented here is to find the sparsest coefficient vector for p, which we call alpha. This is formally known as the basis pursuit denoise problem (BPDN):
compute L1 norm, minimize alpha s.t. (sample - phi*F*alpha) < epsilon
A hack to implement this L1 norm was to iterate on Lasso minimization, tweaking alpha at each iteration. This is wonky but functional...
Next Steps:
At this point in the project the concept of CS-based protein detection has been implemented in prototype form. To move on from here several of the underlying components should be improved, specifically:
The mathematical framework established here for MS/MS samples should also generalize to novel alternative protein sequencing approaches, like the one outlined at the beginning of this project. One concern here though is the issue of sparsity: MS/MS was a nice test case in that the matrices are indeed very sparse, and while RIP was not calculated explicitly here, the technique appears robust. Large numbers of tags or large numbers of fragments may cause issue, however. For these, a Bayesian approach sounds interesting to try. Finally, it will be useful to benchmark this approach against probabilistic protein detection and other current practices.