Vocabulary:
Homolog: sequences in proteins that are the same
pBLAST: "basic logical allignment search tool". Lining up the homologs.
protein family: proteins that share a common eveolutionary origin, reflected by their related functions and similarities in sequences or structure.
primary structure: row of amino acids
secondary structure sequency: write out if it's alpha helix, loop/coil, beta strand. i.e. LLHHHHHLEEEE
secondary structure: alpha helix, loop/coil, beta strand (form beta sheet): found in combination to form the secondary structure of proteins
tertiary structure: "the overall three-dimensional structure resulting from folding and covalent cross-linking of a protein or polynucleotide molecule."
visutalization of proteins: ball and stick, cartoon, surface...
residue type: another name for amino acid
protein hole: a physical hole in a protein. pharmacologists try to identify these holes because they can put drugs in them.
9-mers, 3-mers: 9 amino acids, 3 amino acids.
Part A: Protein analysis
Questions 1.
Q. How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
A. 500g = 3E26 daltons, therefor 500g is ~3E24 amino acids.
Q. Why are there only 20 natural amino acids?
A. Why are there not less? Why are there not more? This is really a discussion point. Having a few more a few less is hard to argue either way, but having an order of magnitude more or less might start to cause problems. For instance, having only 2 amino acids is very little diversity and wouldn't allow the variety of proteins. Having 200 amino acids is a lot of diversity, and might lead to issues and inefficencies. Thinking about this in terms of legos, it's hard to build stuff when you have too many unique pieces, like the super advanced Star Wars ship reconstructions, because you can never find the exact peice you need. It's also hard to build stuff when you have the sets designed for toddlers that are basically all uniform blocks because you can only block structures.
Q. Why most molecular helices are right handed?
A. Folding energy is more favorable... the way the universe formed?
Q. Where did amino acids come from before enzymes that make them, and before life started?
A. Priomordial soup had RNA which could have catelized the creation of amino acids?
Q. What do digital databases and nucleosomes have in common?
A. They both store information.
Questions 2.
Pick any protein (from any organism) of your interest that has a 3D structure and answer the following questions.
Q. Briefly describe the protein you selected and why you selected it.
A. I googled "coolest protein" and found this article. I definately didn't expect the first hit to be deadly proteins, but decided to investigate Major Prion Protein. This is the protein that causes "Mad cow desease" and I found it interesting because we have a very similar protein in our bodies.
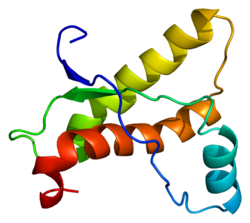
This is what it looks like in cartoon form:
Q. Identity the amino acid sequence of your protein.
A. Found from UniProt.
MANLGCWMLVLFVATWSDLGLCKKRPKPGGWNTGGSRYPGQGSPGGNRYPPQGGGGWGQPHGGGWGQPHGGGWGQPHGGGWGQPHGGGWGQG
GGTHSQWNKPSKPKTNMKHMAGAAAAGAVVGGLGGYMLGSAMSRPIIHFGSDYEDRYYRENMHRYPNQVYYRPMDEYSNQNNFVHDCVNITIKQHT
VTTTTKGENFTETDVKMMERVVEQMCITQYERESQAYYQRGSSMVLFSSPPVILLISFLIFLIVG
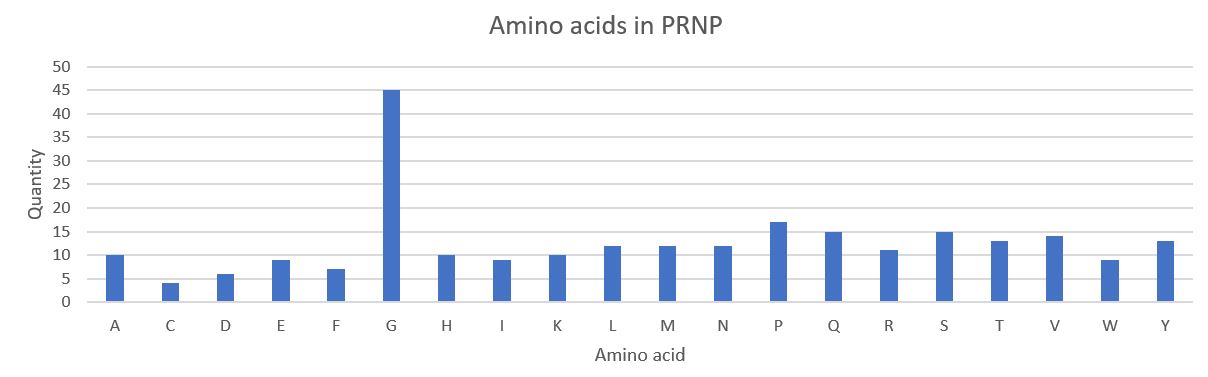
Q. How long is it? What is the most frequent amino acid?
A. 253 amino acids. In comparison see How big is the “average” protein?. The most frequent amino acid in PRNP is G, Glycine.
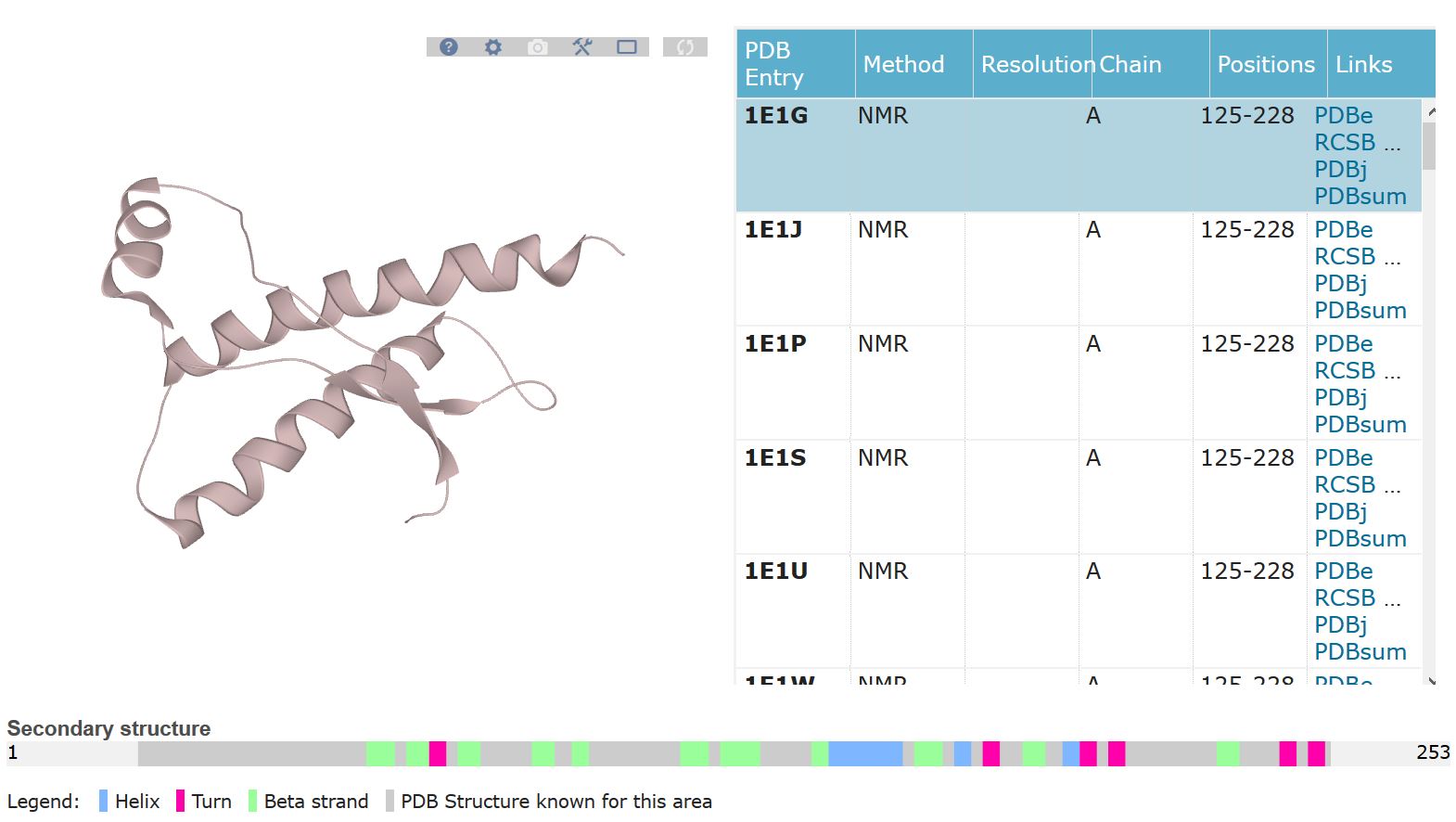
Q. Identify the structure page of your protein in RCSB
A.